69 DNA Repair
Jung Choi; Mary Ann Clark; and Matthew Douglas
Learning Objectives
By the end of this section, you will be able to do the following:
- Discuss the different types of mutations in DNA
- Explain DNA repair mechanisms
DNA replication is a highly accurate process, but mistakes can occasionally occur, such as a DNA polymerase inserting a wrong base. Uncorrected mistakes may sometimes lead to serious consequences, such as cancer. Repair mechanisms correct the mistakes. In rare cases, mistakes are not corrected, leading to mutations; in other cases, repair enzymes are themselves mutated or defective.
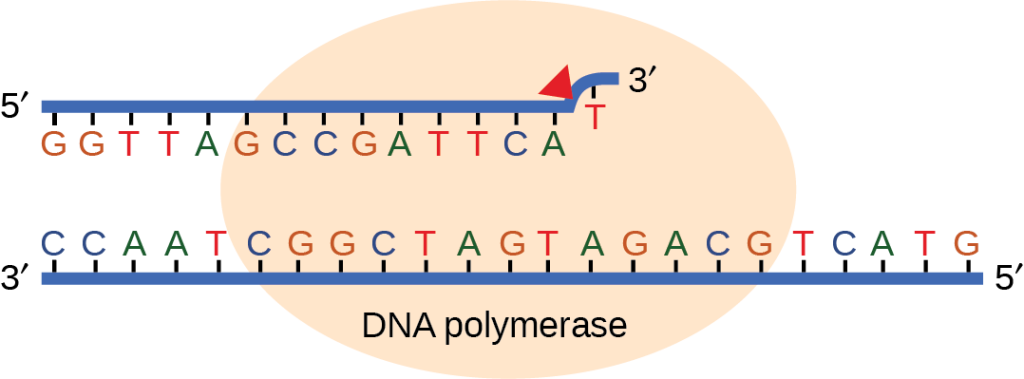
Most of the mistakes during DNA replication are promptly corrected by the proofreading ability of DNA polymerase itself. (Figure 14.17). In proofreading, the DNA pol reads the newly added base before adding the next one, so a correction can be made. The polymerase checks whether the newly added base has paired correctly with the base in the template strand. If it is the right base, the next nucleotide is added. If an incorrect base has been added, the enzyme makes a cut at the phosphodiester bond and releases the wrong nucleotide. This is performed by the 3′ exonuclease action of DNA pol. Once the incorrect nucleotide has been removed, it can be replaced by the correct one.
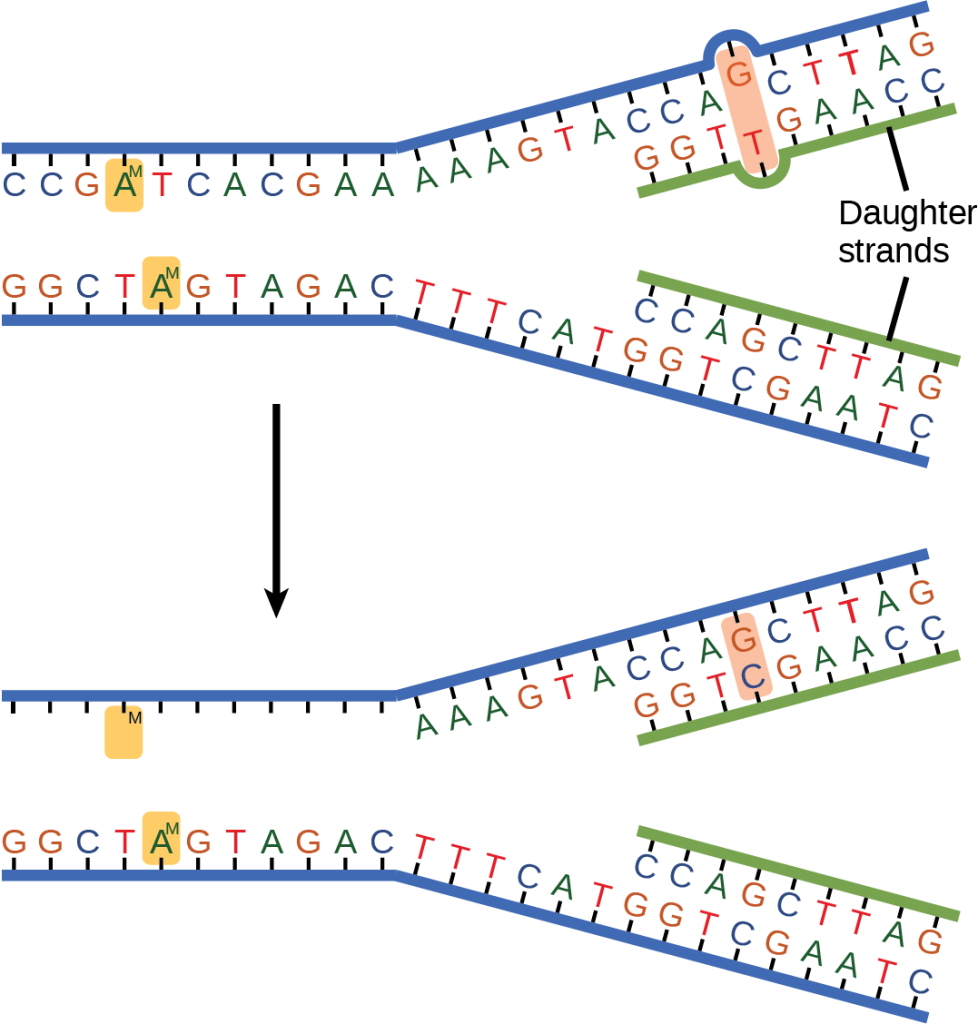
Some errors are not corrected during replication, but are instead corrected after replication is completed; this type of repair is known as mismatch repair (Figure 14.18). Specific repair enzymes recognize the mispaired nucleotide and excise part of the strand that contains it; the excised region is then resynthesized. If the mismatch remains uncorrected, it may lead to more permanent damage when the mismatched DNA is replicated. How do mismatch repair enzymes recognize which of the two bases is the incorrect one? In E. coli, after replication, the nitrogenous base adenine acquires a methyl group; the parental DNA strand will have methyl groups, whereas the newly synthesized strand lacks them. Thus, DNA polymerase is able to remove the wrongly incorporated bases from the newly synthesized, non-methylated strand. In eukaryotes, the mechanism is not very well understood, but it is believed to involve recognition of unsealed nicks in the new strand, as well as a short-term continuing association of some of the replication proteins with the new daughter strand after replication has completed.
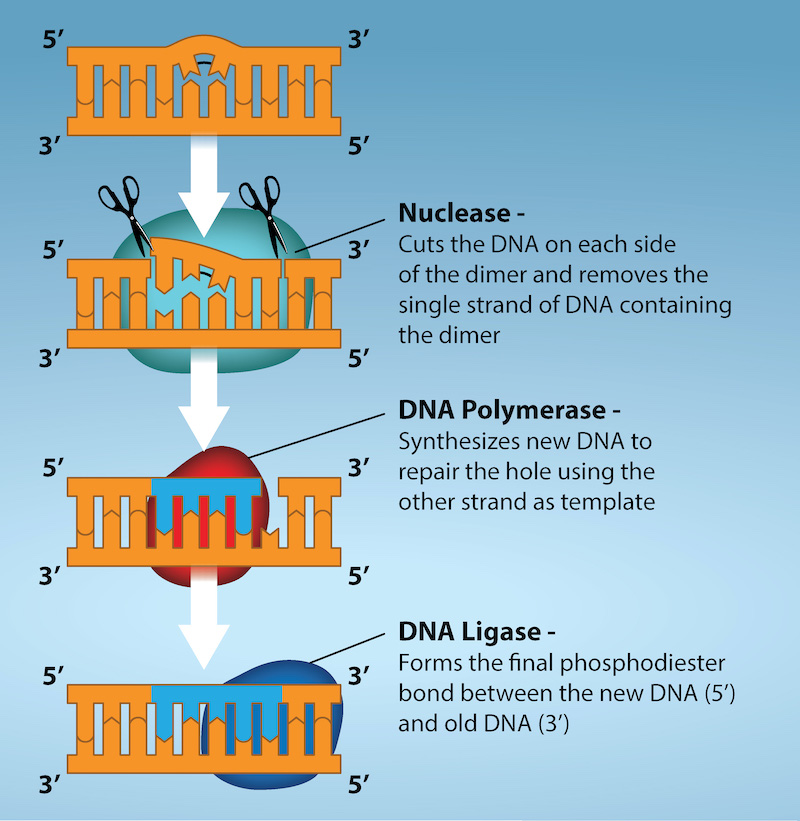
Another type of repair mechanism, nucleotide excision repair, is similar to mismatch repair, except that it is used to remove damaged bases rather than mismatched ones. The repair enzymes replace abnormal bases by making a cut on both the 3′ and 5′ ends of the damaged base (Figure 14.19). The segment of DNA is removed and replaced with the correctly paired nucleotides by the action of DNA pol. Once the bases are filled in, the remaining gap is sealed with a phosphodiester linkage catalyzed by DNA ligase. This repair mechanism is often employed when UV exposure causes the formation of pyrimidine dimers.
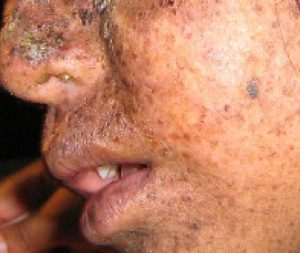
A well-studied example of mistakes not being corrected is seen in people suffering from xeroderma pigmentosa (Figure 14.20). Affected individuals have skin that is highly sensitive to UV rays from the sun. When individuals are exposed to UV light, pyrimidine dimers, especially those of thymine, are formed; people with xeroderma pigmentosa are not able to repair the damage. These are not repaired because of a defect in the nucleotide excision repair enzymes, whereas in normal individuals, the thymine dimers are excised and the defect is corrected. The thymine dimers distort the structure of the DNA double helix, and this may cause problems during DNA replication. People with xeroderma pigmentosa may have a higher risk of contracting skin cancer than those who don’t have the condition.
Mutations in repair genes have been known to cause cancer. Many mutated repair genes have been implicated in certain forms of pancreatic cancer, colon cancer, and colorectal cancer. Mutations can affect either somatic cells or germ cells. If many mutations accumulate in a somatic cell, they may lead to problems such as the uncontrolled cell division observed in cancer. If a mutation takes place in germ cells, the mutation will be passed on to the next generation, as in the case of hemophilia and xeroderma pigmentosa.

